Gaussian processes with outliers
Suppose there are data distributed according a noisy Gaussian process with outliers at places. Doing inference with outliers renders the inference useless, and is indeed where point-wise marginal likelihood maximisation falls flat on its face. With JAXNS we can marginalise over hyper parameters as easily as defining them as prior RVs and marginalising over the posterior.
A Gaussian process is defined by a covariance function, \(K : \mathcal{X} \times \mathcal{X} \to \mathbb{R}\), and a mean function \(\mu : \mathcal{X} \to \mathbb{R}\). Given the above data we see that it is equivalent to a Gaussian likelihood, with Gaussian process prior,
\(L(x) = p(y | x) = \mathcal{N}[y \mid x, \Sigma]\)
and
\(p(x) = \mathcal{N}[x \mid \mu(X), K(X,X)]\)
where \(\mu(X)\) and \(K(X,X)\) are the mean and covariance functions evaluated over the coordinate locations of the data.
The evidence of this model is well known,
\(Z \triangleq p(y) = \int_\mathcal{X} L(x) p(x) \,\mathrm{d} x = \mathcal{N}[y \mid \mu(X), K(X,X) + \Sigma)\)
and likewise the posterior distribution is,
\(p(x \mid y) = \mathcal{N}[x \mid \mu', K']\)
where
\(\mu' = \mu(X) + K(X,X) (K(X,X) + \Sigma)^{-1}(y - \mu(X))\)
and
\(K' = K(X,X) - K(X,X) (K(X,X) + \Sigma)^{-1} K(X,X)\)
Marginalisation
The mean and covariance functions are not a priori known and thus we must infer them as well. Let the hyper parameters of the mean and covariance functions, and the noise covariance be \(\theta\), and suppose we wish to infer their values. The likelihood then becomes,
\(p(y \mid \theta) = \int_\mathcal{X} L(x | \theta) p(x) p(\theta) \,\mathrm{d} x = \mathcal{N}[y \mid \mu_\theta(X), K_\theta(X,X) + \Sigma_\theta)\)
where we recognise this as the marginal likelihood.
Now suppose we wish to predict \(x\) at new points \(X' \subset \mathcal{X}\), then this equivalent to sampling from the marginalised predictive posterior,
Now since \(p(x(X') \mid x(X))\) and \(p(x(X) \mid y, \theta)\) are both Gaussians, their product is also a Gaussian, and is given by,
where
\(m = K(X',X)K(X,X)^{-1} \mu'\)
and
\(S = K(X',X') + K(X',X) (K(X,X)^{-1} K' K(X,X)^{-1} - K(X,X)^{-1})K(X,X')\)
Therefore, sampling from the marginalised predictive distribution is equivalent to sampling \(\theta \sim p(\theta \mid y)\), and then sampling \(x(X') \sim \mathcal{N}[x(X') \mid m, S]\).
[1]:
# For Gaussian processes 64bit is important
from jax.config import config
config.update("jax_enable_x64", True)
import tensorflow_probability.substrates.jax as tfp
tfpd = tfp.distributions
tfpk = tfp.math.psd_kernels
from jaxns import marginalise_dynamic, ForcedIdentifiability, maximum_a_posteriori_point, evaluate_map_estimate
from jax.scipy.linalg import solve_triangular
from jax import random
from jax import numpy as jnp
import pylab as plt
INFO[2023-12-21 13:27:41,688]: Unable to initialize backend 'cuda': module 'jaxlib.xla_extension' has no attribute 'GpuAllocatorConfig'
INFO[2023-12-21 13:27:41,689]: Unable to initialize backend 'rocm': module 'jaxlib.xla_extension' has no attribute 'GpuAllocatorConfig'
INFO[2023-12-21 13:27:41,690]: Unable to initialize backend 'tpu': INTERNAL: Failed to open libtpu.so: libtpu.so: cannot open shared object file: No such file or directory
WARNING[2023-12-21 13:27:41,690]: An NVIDIA GPU may be present on this machine, but a CUDA-enabled jaxlib is not installed. Falling back to cpu.
[2]:
N = 100
M = N // 2
Xstar = jnp.linspace(-2., 2., N)[:, None]
X = Xstar[:M]
true_uncert = 0.2
spectral_params_true = dict(
logits=jnp.asarray([0., 0.]),
locs=jnp.asarray([1. / 1, 1. / 0.5]),
scales=jnp.asarray([1. / 4., 1 / 0.8])
)
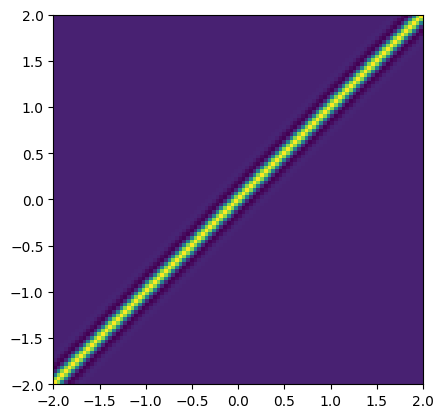
prior_cov = tfpk.SpectralMixture(**spectral_params_true).matrix(Xstar, Xstar) + 1e-12 * jnp.eye(N)
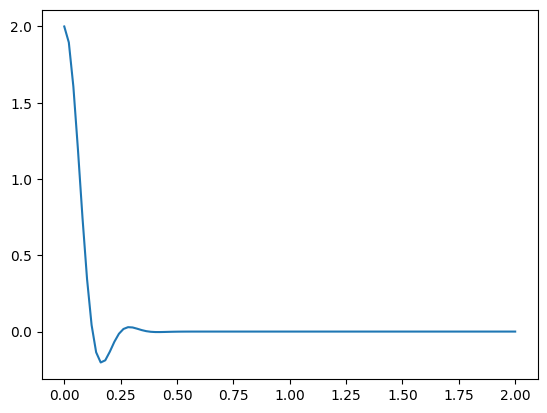
v = jnp.linspace(0., 2., 100)
kern = tfpk.SpectralMixture(**spectral_params_true).matrix(jnp.zeros((1, 1)), v[:, None])
plt.plot(v, kern[0, :])
plt.show()
plt.imshow(prior_cov, origin='lower', extent=(Xstar.min(), Xstar.max(), Xstar.min(), Xstar.max()))
plt.show()
Y = jnp.linalg.cholesky(prior_cov) @ random.normal(random.PRNGKey(42), shape=(N,))
Y_obs = Y[:M] + true_uncert * random.normal(random.PRNGKey(1), shape=(M,))
# Y = jnp.cos(jnp.pi*2. * X[:,0]/2) + jnp.exp(- X[:,0]/2) * jnp.sin(jnp.pi*2. * X[:,0]/3)
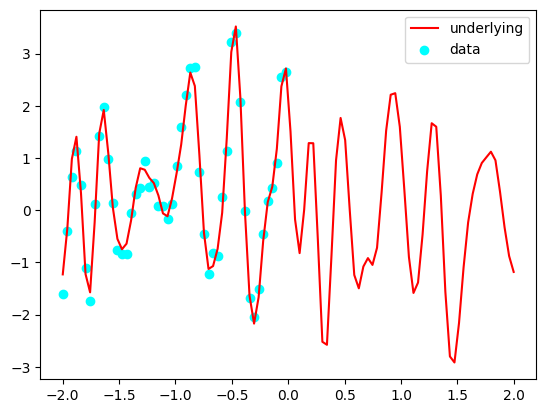
plt.plot(Xstar[:, 0], Y, c='red', label='underlying')
plt.scatter(X[:, 0], Y_obs, c='cyan', label='data')
plt.legend()
plt.show()
[3]:
import jax
from jaxns import Prior, Model, DefaultNestedSampler
kernel = tfpk.SpectralMixture
def log_normal(x, mean, cov):
L = jnp.linalg.cholesky(cov)
# U, S, Vh = jnp.linalg.svd(cov)
log_det = jnp.sum(jnp.log(jnp.diag(L))) # jnp.sum(jnp.log(S))#
dx = x - mean
dx = solve_triangular(L, dx, lower=True)
# U S Vh V 1/S Uh
# pinv = (Vh.T.conj() * jnp.where(S!=0., jnp.reciprocal(S), 0.)) @ U.T.conj()
maha = dx @ dx # dx @ pinv @ dx#solve_triangular(L, dx, lower=True)
log_likelihood = -0.5 * x.size * jnp.log(2. * jnp.pi) - log_det - 0.5 * maha
return log_likelihood
def log_likelihood(uncert, kernel_params):
"""
P(Y|sigma, half_width) = N[Y, f, K]
Args:
uncert: noise
kernel_params: dict of kernel parameters
Returns:
log likelihood
"""
K = tfpk.SpectralMixture(**kernel_params).matrix(X, X)
data_cov = jnp.square(uncert) * jnp.eye(X.shape[0])
mu = jnp.zeros_like(Y_obs)
return log_normal(Y_obs, mu, K + data_cov)
# Build the model
n_components = 2
def prior_model():
smallest_dt = jnp.min(jnp.diff(jnp.sort(X, axis=0), axis=0)) # d
largest_dt = jnp.max(X, axis=0) - jnp.min(X, axis=0) # d
period = yield ForcedIdentifiability(n=2, low=smallest_dt, high=largest_dt, name='period') # n, d
locs = yield Prior(1. / period, name='locs') # n, d
max_bandwidth = (4. * period) # n, d
min_bandwidth = smallest_dt * 2
bandwidth = yield Prior(
tfpd.Uniform(min_bandwidth * jnp.ones_like(max_bandwidth), max_bandwidth * jnp.ones_like(max_bandwidth)),
name='bandwidth') # n, d
scales = yield Prior(1. / bandwidth, name='scales')
logits = yield Prior(tfpd.Normal(0. * jnp.ones(n_components), 1. * jnp.ones(n_components)), name='logits') # n
kernel_params = dict(locs=locs,
scales=scales,
logits=logits)
uncert = yield Prior(tfpd.HalfNormal(0.2), name='uncert')
return uncert, kernel_params
model = Model(prior_model=prior_model, log_likelihood=log_likelihood)
model.sanity_check(random.PRNGKey(0), S=100)
# Create the nested sampler class. In this case without any tuning.
exact_ns = DefaultNestedSampler(
model=model,
max_samples=1e6,
parameter_estimation=True
)
termination_reason, state = jax.jit(exact_ns)(random.PRNGKey(42))
results = exact_ns.to_results(termination_reason=termination_reason, state=state)
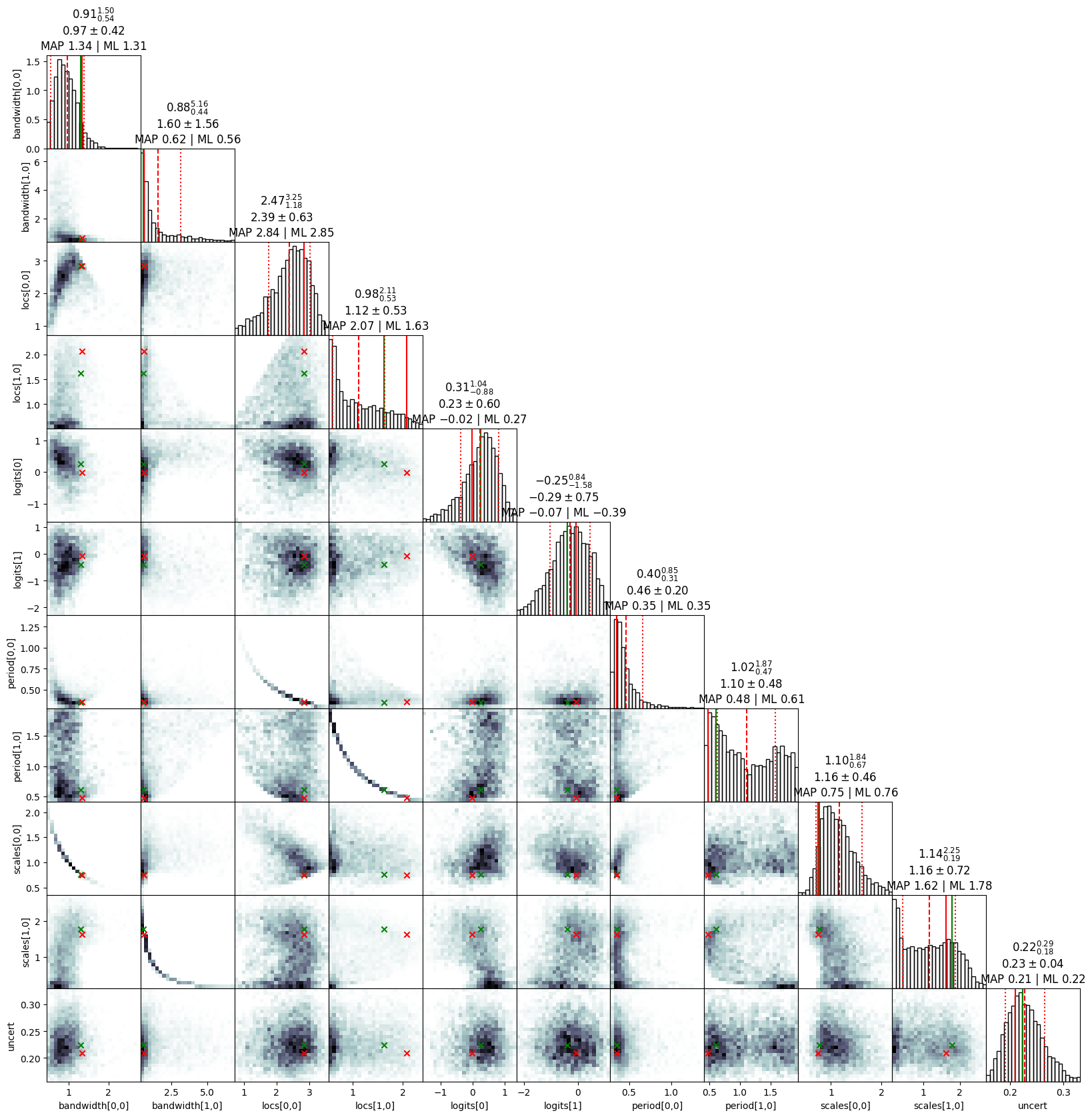
exact_ns.summary(results)
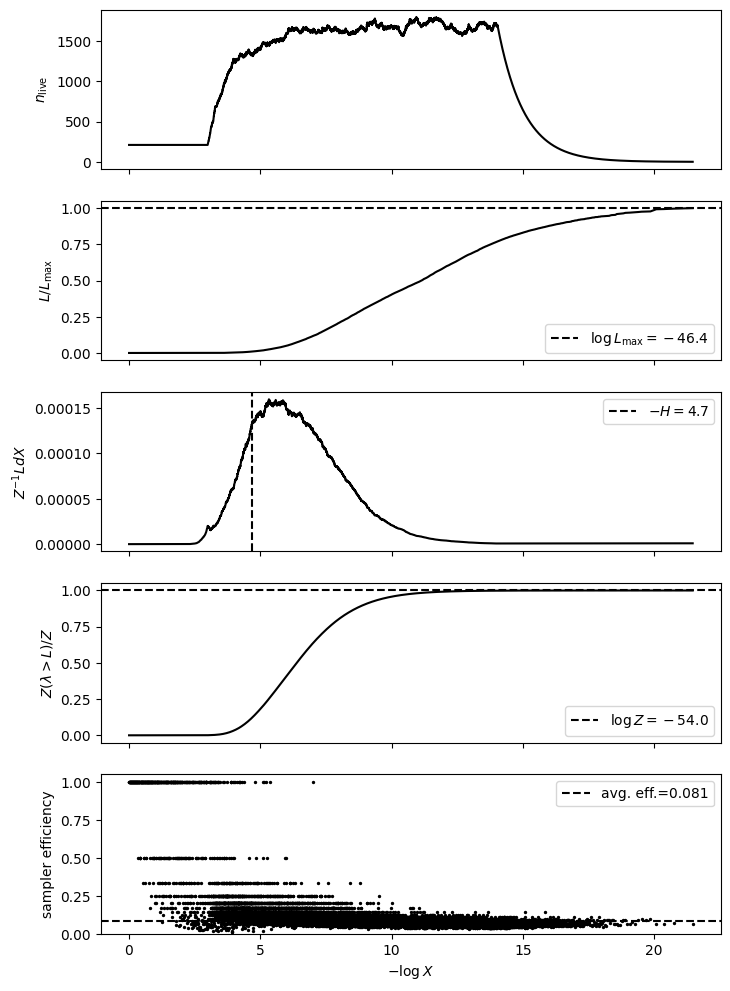
exact_ns.plot_diagnostics(results)
exact_ns.plot_cornerplot(results)
INFO[2023-12-21 13:27:45,178]: Sanity check...
INFO[2023-12-21 13:27:45,434]: Sanity check passed
--------
Termination Conditions:
Small remaining evidence
--------
likelihood evals: 238726
samples: 19320
phantom samples: 16170
likelihood evals / sample: 12.4
phantom fraction (%): 83.7%
--------
logZ=-54.0 +- 0.32
H=-4.69
ESS=720
--------
bandwidth[#]: mean +- std.dev. | 10%ile / 50%ile / 90%ile | MAP est. | max(L) est.
bandwidth[0]: 1.0 +- 0.43 | 0.63 / 0.94 / 1.34 | 1.34 | 1.31
bandwidth[1]: 1.5 +- 1.4 | 0.5 / 0.8 / 3.6 | 0.6 | 0.6
--------
locs[#]: mean +- std.dev. | 10%ile / 50%ile / 90%ile | MAP est. | max(L) est.
locs[0]: 2.42 +- 0.62 | 1.59 / 2.51 / 3.11 | 2.84 | 2.85
locs[1]: 1.12 +- 0.53 | 0.57 / 0.98 / 1.96 | 2.07 | 1.63
--------
logits[#]: mean +- std.dev. | 10%ile / 50%ile / 90%ile | MAP est. | max(L) est.
logits[0]: 0.22 +- 0.59 | -0.52 / 0.3 / 0.85 | -0.02 | 0.27
logits[1]: -0.27 +- 0.72 | -1.19 / -0.22 / 0.58 | -0.07 | -0.39
--------
period[#]: mean +- std.dev. | 10%ile / 50%ile / 90%ile | MAP est. | max(L) est.
period[0]: 0.46 +- 0.2 | 0.32 / 0.4 / 0.63 | 0.35 | 0.35
period[1]: 1.1 +- 0.47 | 0.51 / 1.02 / 1.76 | 0.48 | 0.61
--------
scales[#]: mean +- std.dev. | 10%ile / 50%ile / 90%ile | MAP est. | max(L) est.
scales[0]: 1.13 +- 0.46 | 0.75 / 1.07 / 1.58 | 0.75 | 0.76
scales[1]: 1.21 +- 0.71 | 0.28 / 1.26 / 2.07 | 1.62 | 1.78
--------
uncert: mean +- std.dev. | 10%ile / 50%ile / 90%ile | MAP est. | max(L) est.
uncert: 0.228 +- 0.038 | 0.186 / 0.225 / 0.278 | 0.21 | 0.224
--------
[4]:
def predict_f_fn(uncert, locs, scales, logits):
kernel_params = dict(locs=locs,
scales=scales,
logits=logits)
Kxx = tfpk.SpectralMixture(**kernel_params).matrix(X, X)
Ksx = tfpk.SpectralMixture(**kernel_params).matrix(Xstar, X)
data_cov = jnp.square(uncert) * jnp.eye(X.shape[0])
return Ksx @ jnp.linalg.solve(Kxx + data_cov, Y_obs)
def predict_fvar_fn(uncert, locs, scales, logits):
kernel_params = dict(locs=locs,
scales=scales,
logits=logits)
Kxx = tfpk.SpectralMixture(**kernel_params).matrix(X, X)
Ksx = tfpk.SpectralMixture(**kernel_params).matrix(Xstar, X)
Kss = tfpk.SpectralMixture(**kernel_params).matrix(Xstar, Xstar)
data_cov = jnp.square(uncert) * jnp.eye(X.shape[0])
return jnp.diag(Kss - Ksx @ jnp.linalg.solve(Kxx + data_cov, Ksx.T))
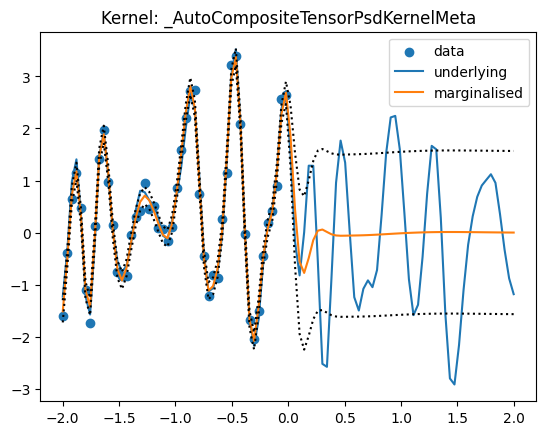
predict_f = marginalise_dynamic(random.PRNGKey(42), results.samples, results.log_dp_mean,
results.ESS, predict_f_fn)
predict_fvar = marginalise_dynamic(random.PRNGKey(42), results.samples, results.log_dp_mean,
results.ESS, predict_fvar_fn)
plt.scatter(X[:, 0], Y_obs, label='data')
plt.plot(Xstar[:, 0], Y, label='underlying')
plt.plot(Xstar[:, 0], predict_f, label='marginalised')
plt.plot(Xstar[:, 0], predict_f + jnp.sqrt(predict_fvar), ls='dotted',
c='black')
plt.plot(Xstar[:, 0], predict_f - jnp.sqrt(predict_fvar), ls='dotted',
c='black')
plt.title("Kernel: {}".format(kernel.__class__.__name__))
plt.legend()
plt.show()
[5]:
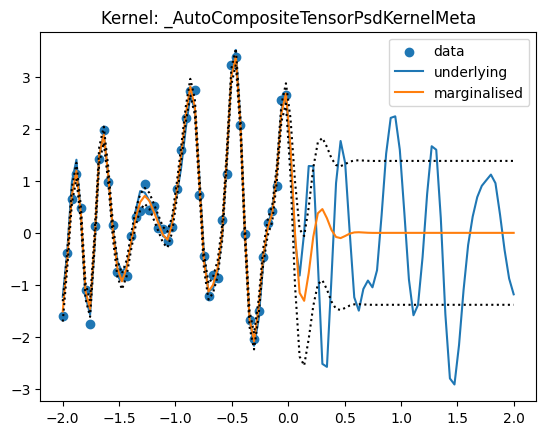
map_sample = maximum_a_posteriori_point(results)
predict_f = evaluate_map_estimate(results, predict_f_fn)
predict_fvar = evaluate_map_estimate(results, predict_fvar_fn)
plt.scatter(X[:, 0], Y_obs, label='data')
plt.plot(Xstar[:, 0], Y, label='underlying')
plt.plot(Xstar[:, 0], predict_f, label='marginalised')
plt.plot(Xstar[:, 0], predict_f + jnp.sqrt(predict_fvar), ls='dotted',
c='black')
plt.plot(Xstar[:, 0], predict_f - jnp.sqrt(predict_fvar), ls='dotted',
c='black')
plt.title("Kernel: {}".format(kernel.__class__.__name__))
plt.legend()
plt.show()
[ ]: