Dual Moons likelihood
Dual moons is a tricky likelihood that mimics physically challenging models.
[1]:
import os
os.environ["XLA_FLAGS"] = "--xla_force_host_platform_device_count=6"
from jax.config import config
config.update("jax_enable_x64", True)
import pylab as plt
import tensorflow_probability.substrates.jax as tfp
from jax import random, numpy as jnp
from jax import vmap
from jaxns import DefaultNestedSampler
from jaxns import Model
from jaxns import Prior
from jaxns import bruteforce_evidence
tfpd = tfp.distributions
INFO[2023-12-21 13:21:14,511]: Unable to initialize backend 'cuda': module 'jaxlib.xla_extension' has no attribute 'GpuAllocatorConfig'
INFO[2023-12-21 13:21:14,511]: Unable to initialize backend 'rocm': module 'jaxlib.xla_extension' has no attribute 'GpuAllocatorConfig'
INFO[2023-12-21 13:21:14,512]: Unable to initialize backend 'tpu': INTERNAL: Failed to open libtpu.so: libtpu.so: cannot open shared object file: No such file or directory
WARNING[2023-12-21 13:21:14,513]: An NVIDIA GPU may be present on this machine, but a CUDA-enabled jaxlib is not installed. Falling back to cpu.
[2]:
from jax._src.scipy.special import logsumexp
def log_likelihood(x):
"""
Dual moon log-likelihood.
"""
def fasym(t):
return jnp.piecewise(
t, [t < 0, t >= 0],
[lambda x: (x / 2) ** 2,
lambda x: (x / 0.5) ** 4
])
xx = x[..., :1]
xy = x[..., 1:2]
rho1 = jnp.sqrt((xx) ** 2 + (xy) ** 2)
theta1 = jnp.arctan2(xy, xx)
term1 = ((rho1 - 2) / 0.05) ** 2
term2 = fasym(theta1 - jnp.pi / 2) / 0.2
rho2 = jnp.sqrt((xx) ** 2 + (xy) ** 2)
theta2 = jnp.arctan2(xy, xx)
term12 = ((rho2 - 2) / 0.05) ** 2
term22 = fasym(theta1 + jnp.pi / 3) / 0.2
pe = logsumexp(-jnp.stack([term1 + term2, term12 + term22]), axis=0)
return pe.squeeze()
def prior_model():
x = yield Prior(tfpd.Uniform(low=-4 * jnp.ones(2), high=4. * jnp.ones(2)), name='x')
return x
model = Model(prior_model=prior_model,
log_likelihood=log_likelihood)
log_Z_true = bruteforce_evidence(model=model, S=250)
print(f"True log(Z)={log_Z_true}")
True log(Z)=-5.104862238439268
[3]:
u_vec = jnp.linspace(0., 1., 250)
args = jnp.stack([x.flatten() for x in jnp.meshgrid(*[u_vec] * model.U_ndims, indexing='ij')], axis=-1)
# The `prepare_func_args(log_likelihood)` turns the log_likelihood into a function that nicely accepts **kwargs
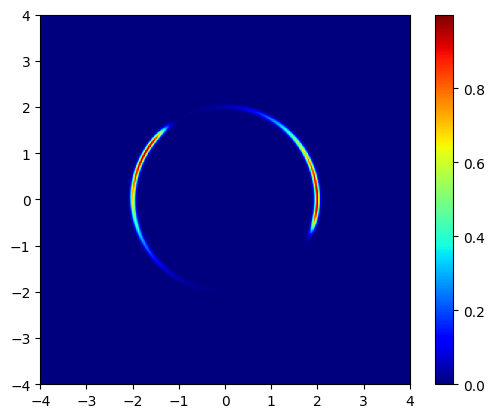
lik = vmap(model.forward)(args).reshape((u_vec.size, u_vec.size))
plt.imshow(jnp.exp(lik), origin='lower', extent=(-4, 4, -4, 4), cmap='jet')
plt.colorbar()
plt.show()
[4]:
# Create the nested sampler class. In this case without any tuning.
ns = DefaultNestedSampler(model=model, max_samples=1e5)
termination_reason, state = ns(random.PRNGKey(42))
results = ns.to_results(termination_reason=termination_reason, state=state)
[5]:
# We can use the summary utility to display results
ns.summary(results)
--------
Termination Conditions:
Small remaining evidence
--------
likelihood evals: 64753
samples: 1680
phantom samples: 720
likelihood evals / sample: 38.5
phantom fraction (%): 42.9%
--------
logZ=-5.07 +- 0.33
H=-4.12
ESS=125
--------
x[#]: mean +- std.dev. | 10%ile / 50%ile / 90%ile | MAP est. | max(L) est.
x[0]: 0.45 +- 0.72 | -0.46 / 0.53 / 1.35 | 1.03 | 1.03
x[1]: 0.0 +- 1.8 | -2.0 / 1.0 / 2.0 | -1.7 | -1.7
--------
[6]:
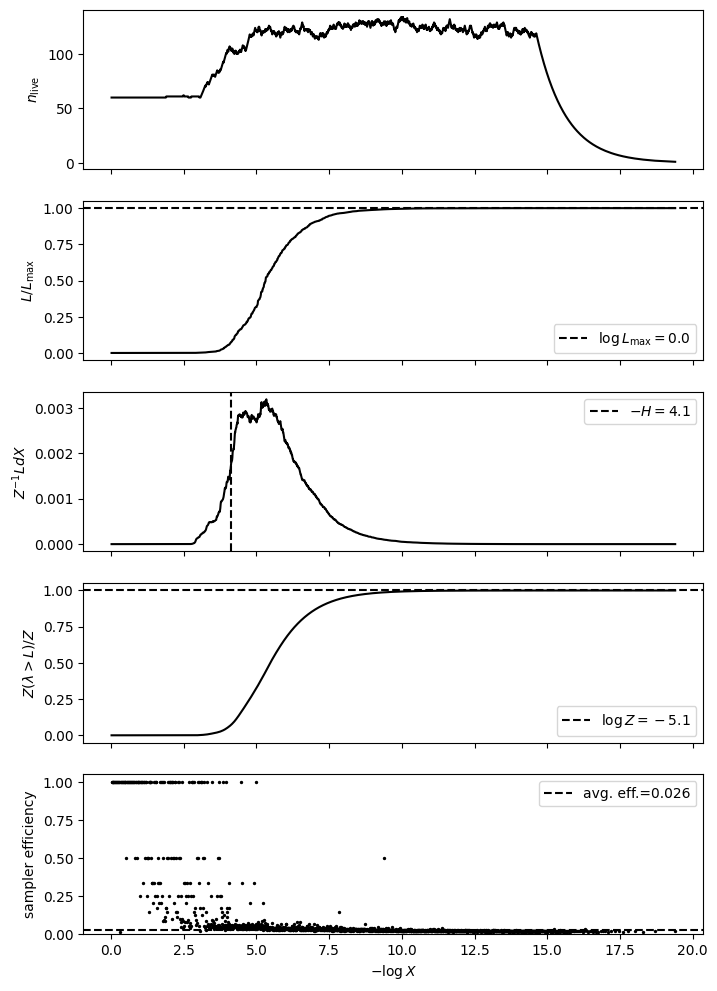
# We plot useful diagnostics and a distribution cornerplot
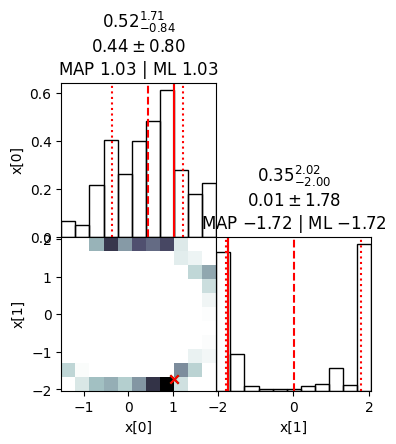
ns.plot_diagnostics(results)
ns.plot_cornerplot(results)