Poisson likelihood and Gamma prior
This is a simple model where our discrete data, \(y\), is modelled as a Poisson RV with Gamma prior, which is a conjugate prior model.
\(L(x) = p(y | x) = \mathcal{P}[y \mid x]\)
and
\(p(x) = \Gamma[x \mid k, \theta]\).
The analytic evidence for this model is,
\(Z = p(y) = \int_\mathcal{X} L(x) p(x) \,\mathrm{d} x = \mathcal{P}[y \mid 1] \frac{\Gamma[1 \mid k, \theta]}{\Gamma[1 \mid k', \theta']}\)
The posterior is also a Gamma distribution,
\(p(x \mid y) = \Gamma[x \mid k', \theta']\)
where
\(k' = k + \sum_i y_i\)
and
\(\theta' = \frac{\theta}{(\theta \sum_i y_i + 1)}\)
[1]:
import numpy as np
import pylab as plt
import tensorflow_probability.substrates.jax as tfp
from jax import random, numpy as jnp
from jaxns import resample
tfpd = tfp.distributions
INFO[2023-12-21 13:26:10,550]: Unable to initialize backend 'cuda': module 'jaxlib.xla_extension' has no attribute 'GpuAllocatorConfig'
INFO[2023-12-21 13:26:10,551]: Unable to initialize backend 'rocm': module 'jaxlib.xla_extension' has no attribute 'GpuAllocatorConfig'
INFO[2023-12-21 13:26:10,552]: Unable to initialize backend 'tpu': INTERNAL: Failed to open libtpu.so: libtpu.so: cannot open shared object file: No such file or directory
WARNING[2023-12-21 13:26:10,552]: An NVIDIA GPU may be present on this machine, but a CUDA-enabled jaxlib is not installed. Falling back to cpu.
[2]:
# Generate data
np.random.seed(42)
num_samples = 10
true_k = 0.5
true_theta = 1.
_gamma = np.random.gamma(true_k, true_theta, size=num_samples)
print(f"Taking {num_samples} samples from a Poisson distribution as data.")
data = jnp.asarray(np.random.poisson(_gamma, size=num_samples))
Taking 10 samples from a Poisson distribution as data.
[3]:
from jaxns import Prior, Model
# Build model
prior_k = 100.
# Note if prior_theta is chosen too large 32-bit will be insufficient
prior_theta = 0.1
def prior_model():
lamda = yield Prior(
tfpd.Gamma(concentration=prior_k, rate=1. / prior_theta),
name='lamda')
return lamda
def log_likelihood(lamda):
"""
Poisson likelihood.
"""
_log_prob = jnp.sum(tfpd.Poisson(rate=lamda).log_prob(data))
return _log_prob
model = Model(prior_model=prior_model, log_likelihood=log_likelihood)
model.sanity_check(random.PRNGKey(0), S=100)
INFO[2023-12-21 13:26:13,013]: Sanity check...
INFO[2023-12-21 13:26:13,410]: Sanity check passed
[4]:
# Evidence and posterior are analytic
def log_gamma_prob(lamda, k, theta):
return tfpd.Gamma(concentration=k, rate=1. / theta).log_prob(lamda)
# return (k-1) * jnp.log(gamma) - gamma / theta - gammaln(k) - k * jnp.log(theta)
true_post_k = prior_k + jnp.sum(data)
true_post_theta = prior_theta / (num_samples * prior_theta + 1.)
true_post_mean_gamma = true_post_theta * true_post_k
true_logZ = log_likelihood(1.) + log_gamma_prob(1., prior_k, prior_theta) - log_gamma_prob(1., true_post_k,
true_post_theta)
print(f"True Evidence = {true_logZ}")
print(f"True posterior concentration (k) = {true_post_k}")
print(f"True posterior rate (1/theta) = {1. / true_post_theta}")
print(f"True posterior lamda = {true_post_mean_gamma}")
True Evidence = -69.31472778320312
True posterior concentration (k) = 100.0
True posterior rate (1/theta) = 20.0
True posterior lamda = 5.0
[5]:
from jaxns import TerminationCondition, DefaultNestedSampler
# Run the nested sampling
ns = DefaultNestedSampler(model=model, num_live_points=100, max_samples=1e4)
term_reason, state = ns(random.PRNGKey(3452345))
results = ns.to_results(termination_reason=term_reason, state=state)
INFO[2023-12-21 13:26:14,390]: Number of parallel Markov-chains set to: 100
[6]:
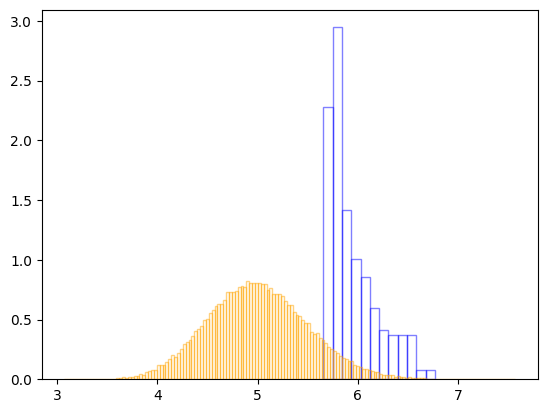
# Comparing samples to true posterior
samples = resample(random.PRNGKey(43083245), results.samples, results.log_dp_mean, S=int(results.ESS))
plt.hist(samples['lamda'], bins='auto', ec='blue', alpha=0.5, density=True, fc='none')
_gamma = np.random.gamma(true_post_k, true_post_theta, size=100000)
plt.hist(_gamma, bins='auto', ec='orange', alpha=0.5, density=True, fc='none')
plt.show()